Battery Cell Design Optimization¶
This example demonstrates how to optimize battery cell design parameters using ionworkspipeline. We optimize active material volume fractions and electrode thicknesses to maximize discharge capacity while preventing lithium plating by constraining the anode potential.
import ionworkspipeline as iwp
from ionworkspipeline.data_fits.models.metrics import Maximum, Minimum, PointBased
from ionworkspipeline.data_fits.models.metrics.actions import (
GreaterThan,
LessThan,
Maximize,
)
import matplotlib.pyplot as plt
import numpy as np
import pybamm
from pybamm import Parameter
/Users/runner/work/ionworks-app/ionworks-app/.venv/lib/python3.12/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Configure DFN Model and Optimization Parameters¶
Import Libraries and Dependencies
# Setup model and baseline parameters
model = pybamm.lithium_ion.DFN()
# Configure symbolic mesh
submesh_types = model.default_submesh_types
for domain in [
"negative electrode",
"separator",
"positive electrode",
"positive particle",
"negative particle",
]:
submesh_types[domain] = pybamm.SymbolicUniform1DSubMesh
# Initialize parameters with thickness constraint
params_baseline = pybamm.ParameterValues("Chen2020")
params_baseline.update({"Cell thickness constraint [m]": 3.045e-3})
# Setup multi-layer cell geometry
N = Parameter("Number of electrodes connected in parallel to make a cell")
# Configure current collectors, electrodes, and separators
params_baseline.update(
{
"Full cell positive current collector thickness [m]": Parameter(
"Positive current collector thickness [m]"
)
* (N + 1),
"Full cell negative current collector thickness [m]": Parameter(
"Negative current collector thickness [m]"
)
* (N + 1),
"Full cell negative electrode thickness [m]": Parameter(
"Negative electrode thickness [m]"
)
* N,
"Full cell positive electrode thickness [m]": Parameter(
"Positive electrode thickness [m]"
)
* N,
"Cell thickness [m]": (
Parameter("Full cell negative electrode thickness [m]")
+ Parameter("Full cell positive electrode thickness [m]")
+ Parameter("Full cell negative current collector thickness [m]")
+ Parameter("Full cell positive current collector thickness [m]")
+ Parameter("Separator thickness [m]") * N
),
}
)
# Link porosity to active material fraction
for electrode in ["Positive", "Negative"]:
params_baseline[f"{electrode} electrode porosity"] = 1.0 - Parameter(
f"{electrode} electrode active material volume fraction"
)
# Define optimization parameters
parameters = {
"Positive electrode active material volume fraction": iwp.Parameter(
"Positive electrode active material volume fraction",
initial_value=0.65,
bounds=(0.5, 0.9),
),
"Negative electrode active material volume fraction": iwp.Parameter(
"Negative electrode active material volume fraction",
initial_value=0.65,
bounds=(0.5, 0.85),
),
"Positive electrode thickness [m]": iwp.Parameter(
"Positive electrode thickness [m]", initial_value=68e-6, bounds=(25e-6, 100e-6)
),
"Negative electrode thickness [m]": iwp.Parameter(
"Negative electrode thickness [m]", initial_value=75e-6, bounds=(30e-6, 105e-6)
),
"Number of electrodes connected in parallel to make a cell": iwp.Parameter(
"Number of electrodes connected in parallel to make a cell",
initial_value=10,
bounds=(1, 50),
),
}
model.variables["Cell thickness [m]"] = pybamm.Parameter("Cell thickness [m]")
print(f"Model configured with {len(parameters)} optimization parameters")
Model configured with 5 optimization parameters
Define Experiment Protocol and Optimization Metrics¶
# Define experiment protocol
experiment = pybamm.Experiment(
[
"Discharge at 60A until 3.0 V",
"Rest for 1 hours",
"Charge at 60A until 4.05 V",
"Charge at 30A until 4.20 V",
"Hold at 4.20 V until C/20",
]
)
# Define optimization actions
actions = {
"Maximize discharge capacity [A.h]": Maximize(
Maximum("Discharge capacity [A.h]"), weight=1.0
),
}
# Constraints define bounds that must be satisfied (using GreaterThan/LessThan actions)
action_constraints = {
"Anode potential minimum constraint [V]": GreaterThan(
Minimum(
"Negative electrode surface potential difference at separator interface [V]"
),
value=0.0,
penalty=150.0,
),
"Cell thickness constraint": LessThan(
PointBased("Cell thickness [m]"),
value=3.045e-3,
),
}
print("Experiment, metrics, and constraints configured")
Experiment, metrics, and constraints configured
Execute Battery Design Optimization¶
# Setup and run optimization
initial_soc = 1.0
params_for_pipeline = {k: v for k, v in params_baseline.items() if k not in parameters}
solver = pybamm.IDAKLUSolver(
options={"max_convergence_failures": 30, "max_error_test_failures": 20}
)
objective_options = {
"model": model,
"simulation_kwargs": {
"experiment": experiment,
"submesh_types": submesh_types,
"solver": solver,
},
}
objective = iwp.objectives.DesignObjective(
options=objective_options,
actions=actions,
constraints=action_constraints,
parameters={"Initial SOC [%]": 100 * initial_soc},
)
optim = iwp.optimizers.AskTellOptimizer(
method="CMAES",
max_iterations=10, # Increase for better results
log_to_screen=True,
)
design_datafit = iwp.DataFit(
objective, parameters=parameters, options={"seed": 0}, optimizer=optim
)
print("Running optimization...")
results = design_datafit.run(params_for_pipeline)
print(f"\nOptimization completed! Final cost: {results.costs:.6f}")
print("Optimal parameters:")
for name in parameters:
print(f" {name}: {results.parameter_values[name]:.6f}")
Running optimization...
Minimising error measure
Using CMAES
Population size: 8
--------------------------------------------------------------------------------
Iter. Eval. Best Current Time m:s
--------------------------------------------------------------------------------
1 9 -3.946107e+01 -3.244782e+01 0:01
2 18 -5.132613e+01 -4.550691e+01 0:03
3 27 -5.984796e+01 -5.069976e+01 0:05
4 36 -6.349323e+01 -6.100279e+01 0:10
5 45 -6.826520e+01 9.999330e+05 0:11
10 90 -6.980606e+01 -6.802096e+01 0:17
--------------------------------------------------------------------------------
Maximum number of iterations (10) reached.
Optimization complete
Optimization completed! Final cost: -69.806059
Optimal parameters:
Positive electrode active material volume fraction: 0.717876
Negative electrode active material volume fraction: 0.740707
Positive electrode thickness [m]: 0.000092
Negative electrode thickness [m]: 0.000094
Number of electrodes connected in parallel to make a cell: 13.387636
Run Validation Simulations¶
# Helper function to extract metrics from solution
def extract_metrics(solution):
sol_discharge = solution.sub_solutions[0]
discharge_capacity = abs(sol_discharge["Discharge capacity [A.h]"]()[-1])
# Combine charge steps
charge_solutions = solution.sub_solutions[2:]
sol_charge = charge_solutions[0]
for charge_sol in charge_solutions[1:]:
sol_charge = sol_charge + charge_sol
anode_potential_series = sol_charge[
"Negative electrode surface potential difference at separator interface [V]"
]()
min_anode_potential = min(anode_potential_series)
mean_anode_potential = anode_potential_series.mean()
return {
"discharge_capacity": discharge_capacity,
"min_anode_potential": min_anode_potential,
"mean_anode_potential": mean_anode_potential,
"solution": solution,
}
# Setup baseline parameters for comparison
params_optimal = params_baseline.copy()
params_optimal.update(results.parameter_values.copy())
params_baseline_original = params_baseline.copy()
for name, param in parameters.items():
params_baseline_original[name] = param.initial_value
# Run simulations for comparison
sim_baseline = iwp.Simulation(
model=model, parameter_values=params_baseline_original, experiment=experiment
)
sim_optimal = iwp.Simulation(
model=model, parameter_values=params_optimal, experiment=experiment
)
sol_baseline = sim_baseline.solve(initial_soc=initial_soc - 0.01)
sol_optimal = sim_optimal.solve(initial_soc=initial_soc - 0.01)
baseline_metrics = extract_metrics(sol_baseline)
optimal_metrics = extract_metrics(sol_optimal)
print("Validation simulations completed")
Validation simulations completed
Analyze Performance Metrics and Parameter Changes¶
# Performance comparison and analysis
capacity_improvement = (
(optimal_metrics["discharge_capacity"] - baseline_metrics["discharge_capacity"])
/ baseline_metrics["discharge_capacity"]
) * 100
min_anode_improvement = (
optimal_metrics["min_anode_potential"] - baseline_metrics["min_anode_potential"]
)
print("Performance Comparison:")
print("=" * 50)
print(f"{'Metric':<35} {'Baseline':<12} {'Optimal':<12} {'Improvement'}")
print("-" * 70)
print(
f"{'Discharge Capacity [A.h]':<35} {baseline_metrics['discharge_capacity']:<12.4f} {optimal_metrics['discharge_capacity']:<12.4f} {capacity_improvement:+.2f}%"
)
print(
f"{'Min Anode Potential [V]':<35} {baseline_metrics['min_anode_potential']:<12.4f} {optimal_metrics['min_anode_potential']:<12.4f} {min_anode_improvement:+.4f}"
)
print(
f"{'Mean Anode Potential [V]':<35} {baseline_metrics['mean_anode_potential']:<12.4f} {optimal_metrics['mean_anode_potential']:<12.4f}"
)
print("=" * 50)
print("\nParameter Changes:")
for name, param in parameters.items():
baseline_val = param.initial_value
optimal_val = results.parameter_values[name]
change = optimal_val - baseline_val
percent_change = (change / baseline_val) * 100
print(f"{name}: {baseline_val:.6f} → {optimal_val:.6f} ({percent_change:+.2f}%)")
# Check thickness constraint
baseline_thickness = (
params_baseline_original.evaluate(params_baseline_original["Cell thickness [m]"])
if "Cell thickness [m]" in params_baseline_original
else "N/A"
)
optimal_thickness = params_optimal.evaluate(params_optimal["Cell thickness [m]"])
print(
f"\nCell Thickness: {baseline_thickness} → {optimal_thickness:.6f} m (limit: {params_baseline['Cell thickness constraint [m]']:.6f} m)"
)
Performance Comparison:
==================================================
Metric Baseline Optimal Improvement
----------------------------------------------------------------------
Discharge Capacity [A.h] 34.9022 69.0529 +97.85%
Min Anode Potential [V] 0.0070 0.0194 +0.0124
Mean Anode Potential [V] 0.0692 0.1016
==================================================
Parameter Changes:
Positive electrode active material volume fraction: 0.650000 → 0.717876 (+10.44%)
Negative electrode active material volume fraction: 0.650000 → 0.740707 (+13.95%)
Positive electrode thickness [m]: 0.000068 → 0.000092 (+34.75%)
Negative electrode thickness [m]: 0.000075 → 0.000094 (+24.86%)
Number of electrodes connected in parallel to make a cell: 10.000000 → 13.387636 (+33.88%)
Cell Thickness: 0.0018579999999999998 → 0.003044 m (limit: 0.003045 m)
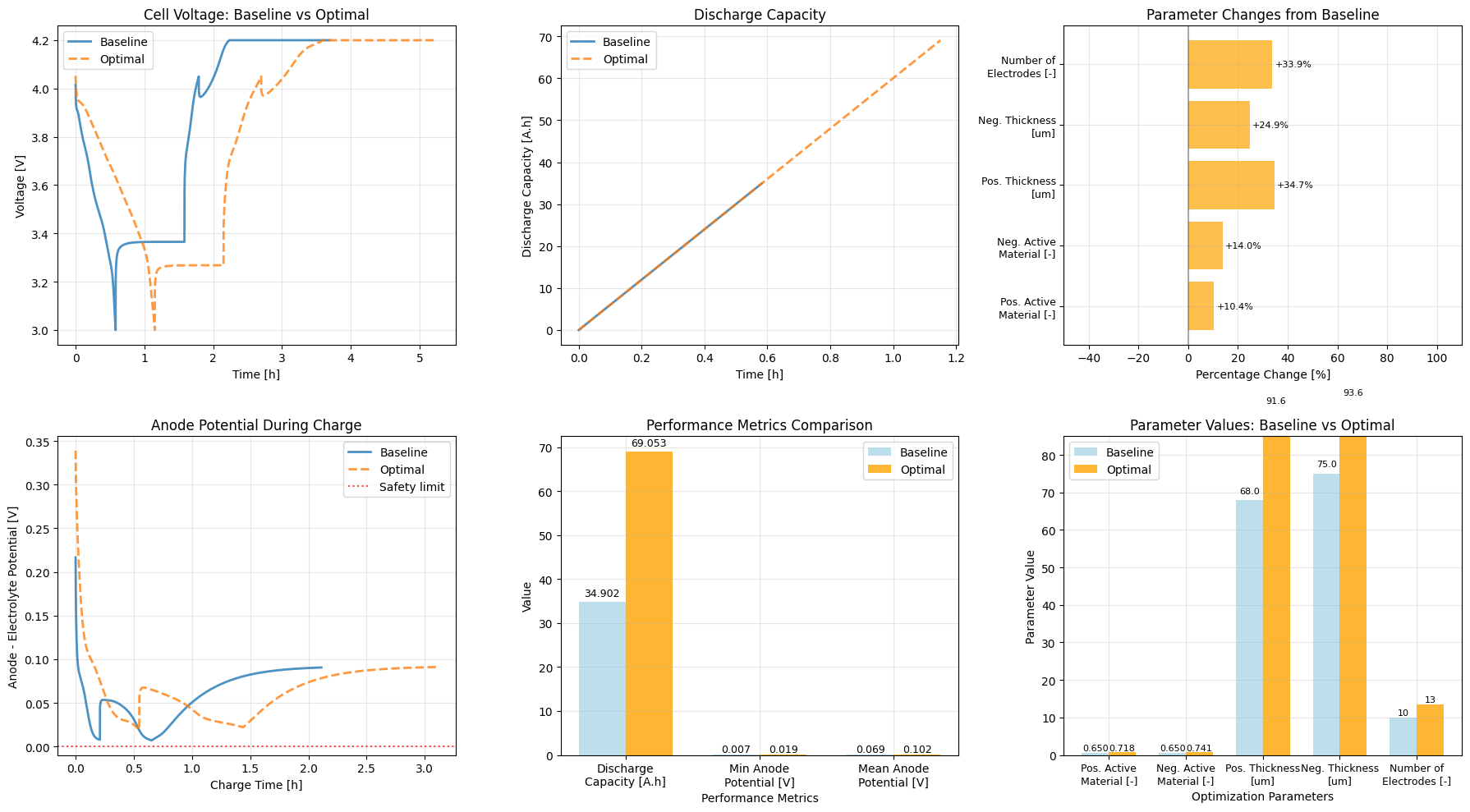
Create Comparative Visualizations¶
# Create comprehensive visualization
fig, axes = plt.subplots(2, 3, figsize=(18, 10))
# Voltage comparison
t_baseline = baseline_metrics["solution"].t / 3600
t_optimal = optimal_metrics["solution"].t / 3600
axes[0, 0].plot(
t_baseline,
baseline_metrics["solution"]["Voltage [V]"](baseline_metrics["solution"].t),
label="Baseline",
linewidth=2,
alpha=0.8,
)
axes[0, 0].plot(
t_optimal,
optimal_metrics["solution"]["Voltage [V]"](optimal_metrics["solution"].t),
label="Optimal",
linewidth=2,
alpha=0.8,
linestyle="--",
)
axes[0, 0].set_xlabel("Time [h]")
axes[0, 0].set_ylabel("Voltage [V]")
axes[0, 0].set_title("Cell Voltage: Baseline vs Optimal")
axes[0, 0].legend()
axes[0, 0].grid(True, alpha=0.3)
# Discharge capacity
sol_discharge_baseline = baseline_metrics["solution"].sub_solutions[0]
sol_discharge_optimal = optimal_metrics["solution"].sub_solutions[0]
axes[0, 1].plot(
sol_discharge_baseline.t / 3600,
abs(sol_discharge_baseline["Discharge capacity [A.h]"](sol_discharge_baseline.t)),
label="Baseline",
linewidth=2,
alpha=0.8,
)
axes[0, 1].plot(
sol_discharge_optimal.t / 3600,
abs(sol_discharge_optimal["Discharge capacity [A.h]"](sol_discharge_optimal.t)),
label="Optimal",
linewidth=2,
alpha=0.8,
linestyle="--",
)
axes[0, 1].set_xlabel("Time [h]")
axes[0, 1].set_ylabel("Discharge Capacity [A.h]")
axes[0, 1].set_title("Discharge Capacity")
axes[0, 1].legend()
axes[0, 1].grid(True, alpha=0.3)
# Anode potential during charge
charge_solutions_baseline = baseline_metrics["solution"].sub_solutions[2:]
charge_solutions_optimal = optimal_metrics["solution"].sub_solutions[2:]
sol_charge_baseline = charge_solutions_baseline[0]
sol_charge_optimal = charge_solutions_optimal[0]
for charge_sol in charge_solutions_baseline[1:]:
sol_charge_baseline = sol_charge_baseline + charge_sol
for charge_sol in charge_solutions_optimal[1:]:
sol_charge_optimal = sol_charge_optimal + charge_sol
t_charge_baseline = (sol_charge_baseline.t / 3600) - (sol_charge_baseline.t[0] / 3600)
t_charge_optimal = (sol_charge_optimal.t / 3600) - (sol_charge_optimal.t[0] / 3600)
anode_baseline = sol_charge_baseline[
"Negative electrode surface potential difference at separator interface [V]"
](sol_charge_baseline.t)
anode_optimal = sol_charge_optimal[
"Negative electrode surface potential difference at separator interface [V]"
](sol_charge_optimal.t)
axes[1, 0].plot(
t_charge_baseline, anode_baseline, label="Baseline", linewidth=2, alpha=0.8
)
axes[1, 0].plot(
t_charge_optimal,
anode_optimal,
label="Optimal",
linewidth=2,
alpha=0.8,
linestyle="--",
)
axes[1, 0].axhline(y=0.0, color="r", linestyle=":", alpha=0.7, label="Safety limit")
axes[1, 0].set_xlabel("Charge Time [h]")
axes[1, 0].set_ylabel("Anode - Electrolyte Potential [V]")
axes[1, 0].set_title("Anode Potential During Charge")
axes[1, 0].legend()
axes[1, 0].grid(True, alpha=0.3)
# Performance metrics bar chart
metrics_names = [
"Discharge\nCapacity [A.h]",
"Min Anode\nPotential [V]",
"Mean Anode\nPotential [V]",
]
baseline_values = [
baseline_metrics["discharge_capacity"],
baseline_metrics["min_anode_potential"],
baseline_metrics["mean_anode_potential"],
]
optimal_values = [
optimal_metrics["discharge_capacity"],
optimal_metrics["min_anode_potential"],
optimal_metrics["mean_anode_potential"],
]
x = np.arange(len(metrics_names))
width = 0.35
axes[1, 1].bar(
x - width / 2,
baseline_values,
width,
label="Baseline",
alpha=0.8,
color="lightblue",
)
axes[1, 1].bar(
x + width / 2, optimal_values, width, label="Optimal", alpha=0.8, color="orange"
)
for i, (baseline, optimal) in enumerate(
zip(baseline_values, optimal_values, strict=False)
):
y_offset = baseline * 0.02 if baseline > 0 else 0.005
axes[1, 1].text(
i - width / 2,
baseline + y_offset,
f"{baseline:.3f}",
ha="center",
va="bottom",
fontsize=9,
)
axes[1, 1].text(
i + width / 2,
optimal + y_offset,
f"{optimal:.3f}",
ha="center",
va="bottom",
fontsize=9,
)
axes[1, 1].set_xlabel("Performance Metrics")
axes[1, 1].set_ylabel("Value")
axes[1, 1].set_title("Performance Metrics Comparison")
axes[1, 1].set_xticks(x)
axes[1, 1].set_xticklabels(metrics_names)
axes[1, 1].legend()
axes[1, 1].grid(True, alpha=0.3)
# Parameter comparison bar chart
param_names = list(parameters.keys())
param_labels = [
"Pos. Active\nMaterial [-]",
"Neg. Active\nMaterial [-]",
"Pos. Thickness\n[um]",
"Neg. Thickness\n[um]",
"Number of\nElectrodes [-]",
]
baseline_param_values = [param.initial_value for param in parameters.values()]
optimal_param_values = [results.parameter_values[name] for name in param_names]
# Convert thickness values to micrometers for better readability
baseline_param_values[2] *= 1e6 # Positive thickness to um
baseline_param_values[3] *= 1e6 # Negative thickness to um
optimal_param_values[2] *= 1e6 # Positive thickness to um
optimal_param_values[3] *= 1e6 # Negative thickness to um
x_params = np.arange(len(param_names))
width_params = 0.35
axes[1, 2].bar(
x_params - width_params / 2,
baseline_param_values,
width_params,
label="Baseline",
alpha=0.8,
color="lightblue",
)
axes[1, 2].bar(
x_params + width_params / 2,
optimal_param_values,
width_params,
label="Optimal",
alpha=0.8,
color="orange",
)
# Add value labels on bars
for i, (baseline, optimal) in enumerate(
zip(baseline_param_values, optimal_param_values, strict=False)
):
y_offset_baseline = baseline * 0.02 if baseline > 0 else 0.005
y_offset_optimal = optimal * 0.02 if optimal > 0 else 0.005
# Format based on parameter type
if i < 2: # Volume fractions
baseline_text = f"{baseline:.3f}"
optimal_text = f"{optimal:.3f}"
elif i < 4: # Thicknesses (now in um)
baseline_text = f"{baseline:.1f}"
optimal_text = f"{optimal:.1f}"
else: # Number of electrodes
baseline_text = f"{baseline:.0f}"
optimal_text = f"{optimal:.0f}"
axes[1, 2].text(
i - width_params / 2,
baseline + y_offset_baseline,
baseline_text,
ha="center",
va="bottom",
fontsize=8,
)
axes[1, 2].text(
i + width_params / 2,
optimal + y_offset_optimal,
optimal_text,
ha="center",
va="bottom",
fontsize=8,
)
axes[1, 2].set_xlabel("Optimization Parameters")
axes[1, 2].set_ylabel("Parameter Value")
axes[1, 2].set_title("Parameter Values: Baseline vs Optimal")
axes[1, 2].set_xticks(x_params)
axes[1, 2].set_ylim(top=85)
axes[1, 2].set_xticklabels(param_labels, fontsize=9)
axes[1, 2].legend()
axes[1, 2].grid(True, alpha=0.3)
# Add upper right subplot for summary statistics
param_changes = []
for _i, (name, param) in enumerate(parameters.items()):
baseline_val = param.initial_value
optimal_val = results.parameter_values[name]
percent_change = ((optimal_val - baseline_val) / baseline_val) * 100
param_changes.append(percent_change)
axes[0, 2].barh(
range(len(param_names)),
param_changes,
color=["orange" if x > 0 else "lightblue" for x in param_changes],
alpha=0.7,
)
axes[0, 2].set_yticks(range(len(param_names)))
axes[0, 2].set_yticklabels(param_labels, fontsize=9)
axes[0, 2].set_xlabel("Percentage Change [%]")
axes[0, 2].set_title("Parameter Changes from Baseline")
axes[0, 2].set_xlim(-50, 110)
axes[0, 2].axvline(x=0, color="black", linestyle="-", alpha=0.3)
axes[0, 2].grid(True, alpha=0.3)
# Add percentage labels
for i, change in enumerate(param_changes):
axes[0, 2].text(
change + (1 if change >= 0 else -1),
i,
f"{change:+.1f}%",
va="center",
ha="left" if change >= 0 else "right",
fontsize=8,
)
plt.tight_layout()
plt.show()
print("Optimization analysis completed!")
Optimization analysis completed!