1. Half-cell OCP¶
In this example, we use synthetic GITT data to fit the MSMR model of the half-cell open-circuit potential (OCP) for a graphite electrode.
Before running this example, make sure to run the script true_parameters/generate_data.py to generate the synthetic data.
from pathlib import Path
import ionworkspipeline as iwp
import iwutil
/Users/runner/work/ionworks-app/ionworks-app/.venv/lib/python3.12/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
MSMR model theory¶
Here we briefly outline the MSMR model, as described in Baker and Verbrugge (2018).
The MSMR model is developed by assuming that all electrochemical reactions at the electrode/electrolyte interface in a lithium insertion cell can be expressed in the form
The equation for each reaction can be inverted to give
At a particle interface with the electrolyte, local equilibrium requires that
Load the data¶
We load the synthetic GITT data for the negative electrode and the extract the OCP from the relaxed voltages.
gitt_data = iwutil.read_df(
Path("synthetic_data") / "half_cell_negative_electrode" / "gitt.csv"
)
ocp_data = iwp.data_fits.preprocess.pulse_data_to_ocp(gitt_data)
Set up the parameters to fit¶
We first set up our initial guesses for the parameters. For the MSMR model, good initial guesses can be found manually by looking at the data (e.g. dVdQ analysis) or by using the built-in function iwp.data_fits.objectives.msmr_half_cell_initial_guess.
Let’s firs set up some initial guesses manually to get a feel for how the parameter values affect the model. We need to specify initial values for \(U_j\), \(X_j\), and \(\omega_j\), as well as the total capacity of the electrode. We can also specify a “lower excess capacity” parameter, which is the amount of capacity that is not accessed in the experiment at the lower end of the stoichiometry. In practice, the electrode will no be fully delithiated at the start of the experiment - this parameter describes the amount of capacity required to fully delithiate the electrode from its initial state in the experiment.
We can use the iwp.data_fits.objectives.plot_each_species_msmr function to help with this. This function plots the OCP and dQ/dU for each reaction. Each reaction has a corresponding peak in the dQ/dU curve, so we can use this to help us pick reasonable initial values.
Try changing the initial values and re-running the cell to see how the model changes.
Q_use = ocp_data["Capacity [A.h]"].max()
param_init = {
"Negative electrode host site standard potential [V]": [
0.08,
0.13,
0.15,
0.22,
0.22,
0.55,
],
"Negative electrode host site occupancy fraction": [
0.30,
0.25,
0.15,
0.11,
0.03,
0.13,
],
"Negative electrode host site ideality factor": [0.1, 0.04, 0.9, 4.1, 0.05, 8.5],
"Negative electrode capacity [A.h]": Q_use,
"Negative electrode lower excess capacity [A.h]": 0.01 * Q_use,
}
fig, ax = iwp.data_fits.objectives.plot_each_species_msmr(
ocp_data, param_init, "negative", voltage_limits=(0, 1.5)
)
ax[1].set_xlim(0, 50)
(0.0, 50.0)
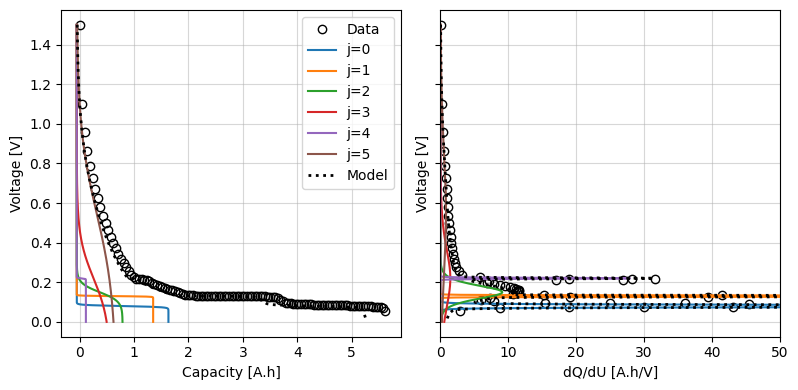
Next we use these initial guesses to set up the parameters for the fit. Each of the fit parameters must be an iwp.Parameter object, which has a name, initial value, and bounds. We can create these objects manually, for example:
a = iwp.Parameter("a", initial_value=1.0, bounds=(0, 2))
but for the MSMR model, we can use the iwp.data_fits.objectives.get_msmr_params_for_fit function to automatically create these objects. The function takes the electrode name and initial guesses, and returns a dictionary of iwp.Parameter objects. Additional keyword arguments, such as a bounds_function that specifies how to set the bounds for the parameters, can be passed to the function to give more control over the fitting process.
Here we use the “Qj” format, where the capacity of each reaction is a fit parameter, rather than the fractional site occupancies \(X_j\). Note this function does not add the lower excess capacity parameter, so we need to do this separately.
parameters = iwp.data_fits.objectives.get_msmr_params_for_fit(
param_init, "negative", Q_use, species_format="Qj"
)
# Add the lower excess capacity parameter
parameters.update(
{
"Negative electrode lower excess capacity [A.h]": (
iwp.Parameter(
"Q_lowex", initial_value=0.01 * Q_use, bounds=(0, 0.2 * Q_use)
)
)
}
)
Run the fit¶
Now we are ready to run the fit. We create the objective, the DataFit object, and run the fit.
First, we create the MSMRHalfCell objective function and pass in the data. We can also specify any additional options, such as the model to use and callbacks to interact with the fitting process (e.g. to configure custom plots or logging). Here we also pass a “dUdQ cutoff” option, that tells the objective to ignore the data above a certain dU/dQ value. We use the calculate_dUdQ_cutoff function to calculate this value from the data, based on some reasonable default settings.
Next, we create the DataFit object and pass in the objective and the parameters to fit. We can also customize the fitting process by passing in a cost function and selecting an optimizer.
Here, the only known parameter is the ambient temperature, which we set to 25°C. This should be the temperature at which the experiment was conducted.
known_params = {"Ambient temperature [K]": 298.15}
# Create the objective
dUdQ_cutoff = iwp.data_fits.util.calculate_dUdQ_cutoff(ocp_data)
objective = iwp.objectives.MSMRHalfCell(
ocp_data,
options={
"dUdQ cutoff": dUdQ_cutoff,
"model": iwp.data_fits.models.MSMRHalfCellModel("negative"),
},
)
# Create the DataFit object and run the fit
optimizer = iwp.optimizers.ScipyLeastSquares()
datafit = iwp.DataFit(
objective,
parameters=parameters,
optimizer=optimizer,
multistarts=10,
options={"seed": 0},
)
results = datafit.run(known_params)
/var/folders/p1/6r4c82w967v9djw141qtq0vm0000gn/T/ipykernel_30245/488006838.py:4: DeprecationWarning: calculate_dUdQ_cutoff is deprecated. Use data.calculate_dUdQ_cutoff() on a DataLoader instance instead.
dUdQ_cutoff = iwp.data_fits.util.calculate_dUdQ_cutoff(ocp_data)
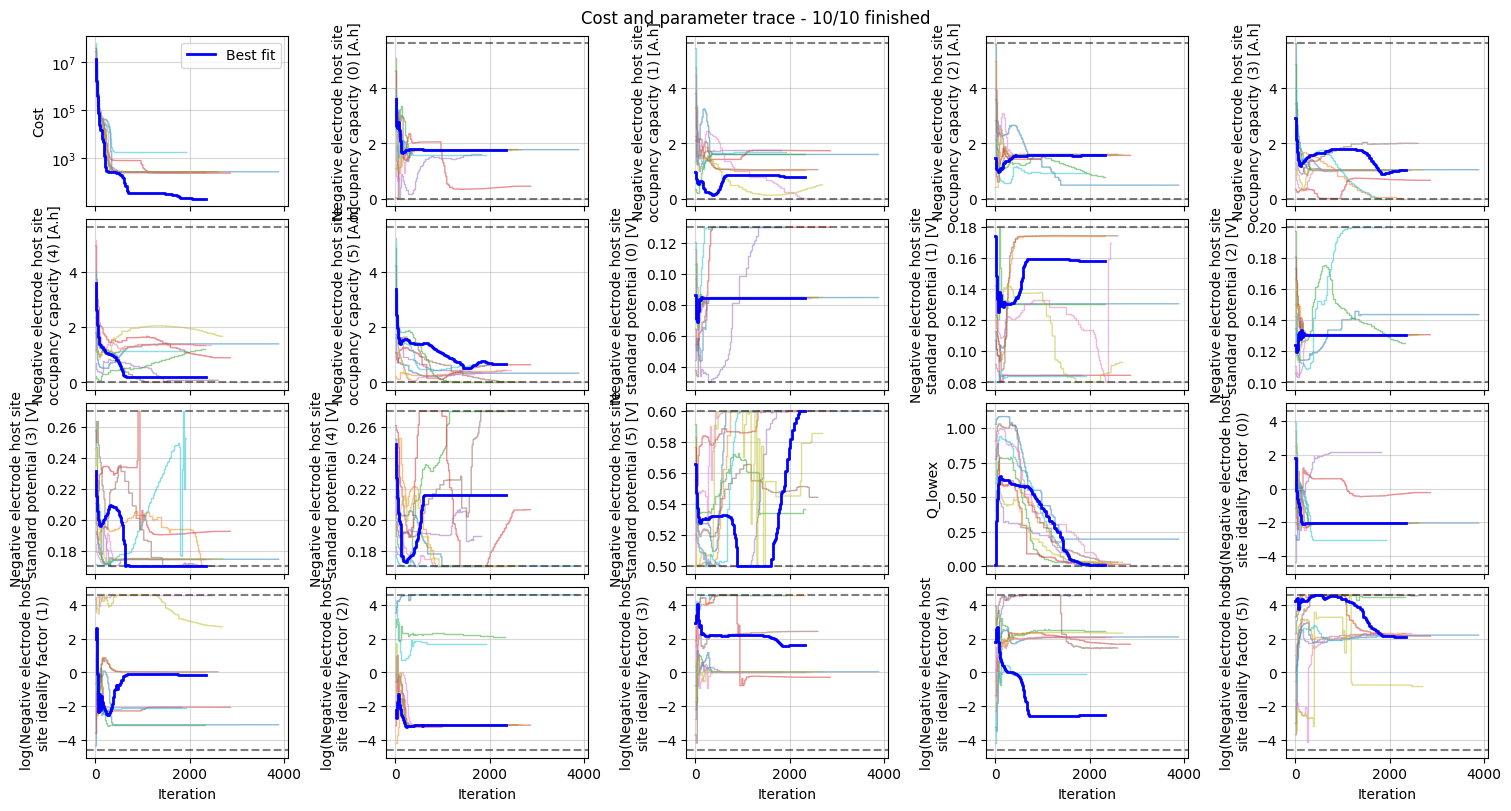
We can plot the trace of the cost function and parameter values during the fit, and the fitted OCP, dU/dQ and dQ/dU curves.
_ = datafit.plot_trace()
_ = datafit.plot_fit_results()
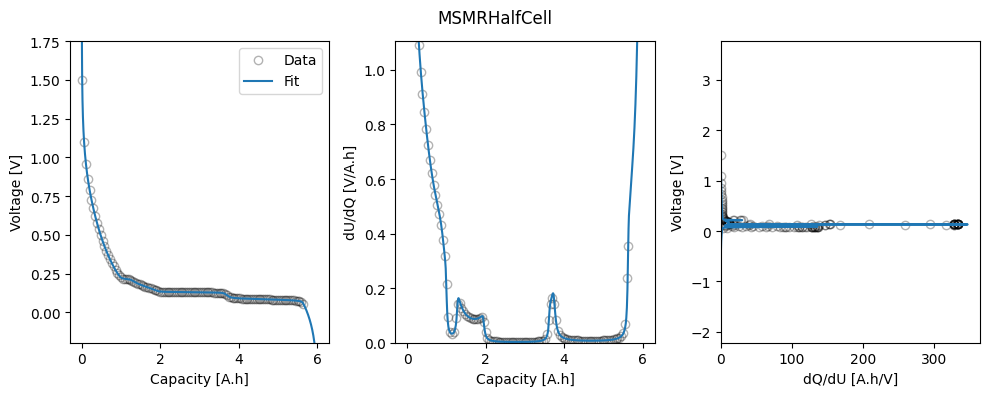
We can take a look at the fitted parameters by accessing the parameter_values attribute of the results object.
results.parameter_values
{'Negative electrode host site standard potential [V]': [0.08463816509796918,
0.15825366905163807,
0.1306570151711749,
0.17000000000010507,
0.21563578955079762,
0.5999999999999177],
'Negative electrode host site ideality factor': [0.12906300951657462,
0.8379022678650083,
0.04336414438996676,
5.152444798412893,
0.07971280133523803,
8.312282354582386],
'Negative electrode host site occupancy capacity [A.h]': [1.7634877181659274,
0.7977751946987168,
1.5610013934968476,
1.0203206681184394,
0.21126251560059772,
0.6506104645154457],
'Negative electrode host site occupancy fraction': [0.29369640548754383,
0.1328638156401908,
0.2599737403943373,
0.1699271900700809,
0.03518427761474983,
0.10835457079309728],
'Negative electrode capacity [A.h]': 6.004457954595975,
'Negative electrode lower excess capacity [A.h]': 0.010101426152755158}