Fitting electrolyte conductivity (Landesfeind 2019 form)¶
Landesfeind & Gasteiger (2019) report measurements of the four binary-electrolyte transport properties — ionic conductivity \(\kappa\), salt diffusion coefficient \(D_e\), thermodynamic factor \(\chi\) and cation transference number \(t_+^0\) — for three solvent systems (EC:DMC (1:1), EC:EMC (3:7), EMC:FEC (19:1)) over a wide \((c_e, T)\) range, and provide a closed-form fit for each property. For the ionic conductivity (in mS/cm, with \(c\) in mol/L and \(T\) in K) the closed form is
with six fitted coefficients \(p_1, \dots, p_6\) per solvent system. (The diffusivity, TDF and transference number have analogous closed forms with their own coefficients — 4, 9 and 9 respectively.)
iwp.direct_entries.landesfeind_electrolyte(c_e, system) packages all four functions plus the published coefficients for the chosen system as a single DirectEntry. Critically, the coefficients are stored as pybamm.Parameters rather than hard-coded numbers, which means we can override any of them with iwp.Parameter in a pipeline to refit them to our own data — without reimplementing the closed form. This notebook demonstrates that workflow on the conductivity coefficients.
We work at a single isotherm (\(T = 293.15\) K, the closest measurement temperature in Landesfeind 2019 to \(25\) °C). We compare three things:
Digitized paper data at \(T = 293.15\) K, plotted against the published equation (no fit — the published \(p_1\dots p_6\) are already in the direct entry).
Mock data generated from the published equation at \(T = 293.15\) K with added Gaussian noise.
A fit to the mock data, recovering a subset of the published coefficients.
At a single \(T\) the six coefficients are not all separately identifiable: \(p_2\) enters only via the prefactor \(A(T) = p_1\,(1 + (T - p_2))\) and is perfectly correlated with \(p_1\) when \(T\) is fixed; likewise the temperature-coupled terms \(p_5\,e^{1000/T}\) and \(p_6\,e^{1000/T}\) collapse to constants. We therefore hold \(p_2\), \(p_5\), \(p_6\) at their published values and fit the remaining three (\(p_1\), \(p_3\), \(p_4\)).
import ionworkspipeline as iwp
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pybamm
/Users/runner/work/ionworks-app/ionworks-app/.venv/lib/python3.12/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
1. The Landesfeind direct entry¶
landesfeind_electrolyte returns a DirectEntry whose parameters dict contains the conductivity, diffusivity, TDF and transference number functions, plus the published numeric values for every coefficient those functions reference. We grab the conductivity function — which still has \(p_1, \dots, p_6\) as pybamm.Parameters — so we can evaluate it later with the published coefficients (for reference curves) and inside the fitting objective (with \(p_1, p_3, p_4\) replaced by InputParameters the optimizer sets).
electrolyte_entry = iwp.direct_entries.landesfeind_electrolyte(
c_e=1000, system="EC:EMC (3:7)"
)
kappa_func = electrolyte_entry.parameters["Electrolyte conductivity [S.m-1]"]
2. Digitized paper data at \(T = 293.15\) K¶
The CSV landesfeind_2019_ec_emc_3_7.csv was sourced from the CALiSol-23 dataset, which digitized the conductivity points from the figures of Landesfeind & Gasteiger 2019. We filter to the \(T = 293.15\) K isotherm.
all_paper_data = pd.read_csv("landesfeind_2019_ec_emc_3_7.csv")
T_iso = 293.15 # K
paper_data = all_paper_data[all_paper_data["Temperature [K]"] == T_iso].reset_index(
drop=True
)
paper_data
| Electrolyte concentration [mol.m-3] | Temperature [K] | Electrolyte conductivity [S.m-1] | |
|---|---|---|---|
| 0 | 102.0 | 293.15 | 0.2467 |
| 1 | 497.0 | 293.15 | 0.6796 |
| 2 | 998.0 | 293.15 | 0.8267 |
| 3 | 1499.0 | 293.15 | 0.7172 |
| 4 | 2003.0 | 293.15 | 0.5047 |
| 5 | 3000.0 | 293.15 | 0.2216 |
3. Generate mock data¶
We sample the published equation on a denser concentration grid at the same temperature and add 5% Gaussian noise. This is what a clean experimental campaign at this isotherm might produce.
rng = np.random.default_rng(0)
c_mock = np.linspace(100, 3000, 12) # mol/m^3
T_mock = np.full_like(c_mock, T_iso)
c_vec = pybamm.Vector(c_mock[:, np.newaxis])
T_vec = pybamm.Vector(T_mock[:, np.newaxis])
kappa_published = (
iwp.ParameterValues(electrolyte_entry.parameters)
.process_symbol(kappa_func(c_vec, T_vec))
.evaluate()
.flatten()
)
kappa_mock = kappa_published * (1 + 0.05 * rng.standard_normal(c_mock.size))
mock_data = pd.DataFrame(
{
"Electrolyte concentration [mol.m-3]": c_mock,
"Temperature [K]": T_mock,
"Electrolyte conductivity [S.m-1]": kappa_mock,
}
)
mock_data.head()
| Electrolyte concentration [mol.m-3] | Temperature [K] | Electrolyte conductivity [S.m-1] | |
|---|---|---|---|
| 0 | 100.000000 | 293.15 | 0.241467 |
| 1 | 363.636364 | 293.15 | 0.591138 |
| 2 | 627.272727 | 293.15 | 0.792801 |
| 3 | 890.909091 | 293.15 | 0.839785 |
| 4 | 1154.545455 | 293.15 | 0.804313 |
4. A custom fitting objective¶
Because we are fitting an algebraic relationship rather than a PyBaMM simulation, we subclass FittingObjective and skip the simulation in build. The conductivity function is taken straight from all_parameter_values (which the pipeline populates from the direct entry) — we don’t reimplement the equation. We bake the data points \((c_e, T)\) into the symbol as pybamm.Vectors once in build, so each optimizer step is a single vectorized evaluate rather than a Python loop. The fitted coefficients \(p_1, p_3, p_4\) remain InputParameters that the optimizer feeds in via the inputs dict.
class LandesfeindConductivityObjective(iwp.objectives.FittingObjective):
def process_data(self):
data = self.data["data"]
self._processed_data = {
"Electrolyte conductivity [S.m-1]": data[
"Electrolyte conductivity [S.m-1]"
].values
}
def build(self, all_parameter_values):
data = self.data["data"]
c_e = data["Electrolyte concentration [mol.m-3]"].values[:, np.newaxis]
T = data["Temperature [K]"].values[:, np.newaxis]
kappa = all_parameter_values["Electrolyte conductivity [S.m-1]"]
self._processed_kappa = all_parameter_values.process_symbol(
kappa(pybamm.Vector(c_e), pybamm.Vector(T))
)
def run(self, inputs, full_output=False):
return {
"Electrolyte conductivity [S.m-1]": self._processed_kappa.evaluate(
inputs=inputs
).flatten()
}
5. Fit \(p_1\), \(p_3\), \(p_4\) to the mock data¶
The fit parameters use the same names as the coefficients defined inside landesfeind_electrolyte, so the pipeline replaces them with InputParameters while leaving \(p_2\), \(p_5\), \(p_6\) at the published values from the direct entry.
fit_parameters = {
name: iwp.Parameter(name, initial_value=v0, bounds=bnds)
for name, v0, bnds in [
("Landesfeind electrolyte conductivity p1", 0.3, (1e-3, 5)),
("Landesfeind electrolyte conductivity p3", -0.5, (-5, 5)),
("Landesfeind electrolyte conductivity p4", 0.5, (-2, 2)),
]
}
objective = LandesfeindConductivityObjective(mock_data)
datafit = iwp.DataFit(objective, parameters=fit_parameters)
pipeline = iwp.Pipeline(
{
"electrolyte": electrolyte_entry,
"fit": datafit,
}
)
fitted_values = pipeline.run()
Compare fitted vs published values:
comparison = pd.DataFrame(
{
"published": [electrolyte_entry.parameters[name] for name in fit_parameters],
"fit (mock data)": [fitted_values[name] for name in fit_parameters],
},
index=list(fit_parameters),
)
comparison
| published | fit (mock data) | |
|---|---|---|
| Landesfeind electrolyte conductivity p1 | 0.521 | 0.511212 |
| Landesfeind electrolyte conductivity p3 | -1.060 | -1.038286 |
| Landesfeind electrolyte conductivity p4 | 0.353 | 0.338020 |
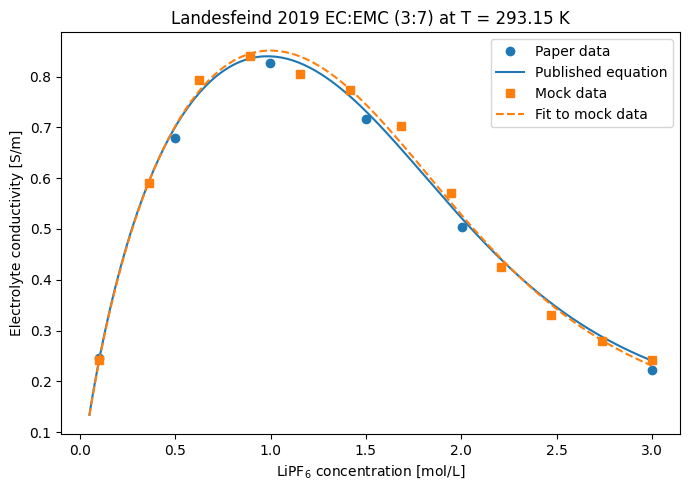
6. Plot at \(T = 293.15\) K¶
We plot, on the same axes:
The 6 digitized paper data points and the published equation curve (no fit — taken straight from the direct entry).
The 12 mock data points and the curve produced by the fit (\(p_1\), \(p_3\), \(p_4\) from the optimizer; \(p_2\), \(p_5\), \(p_6\) from the direct entry’s published values).
c_plot = np.linspace(50, 3000, 200)
c_vec = pybamm.Vector(c_plot[:, np.newaxis])
T_vec = pybamm.Vector(np.full_like(c_plot, T_iso)[:, np.newaxis])
sym = kappa_func(c_vec, T_vec)
published_curve = (
iwp.ParameterValues(electrolyte_entry.parameters)
.process_symbol(sym)
.evaluate()
.flatten()
)
fitted_curve = fitted_values.process_symbol(sym).evaluate().flatten()
fig, ax = plt.subplots(figsize=(7, 5))
ax.plot(
paper_data["Electrolyte concentration [mol.m-3]"] / 1000,
paper_data["Electrolyte conductivity [S.m-1]"],
"o",
color="C0",
label="Paper data",
)
ax.plot(c_plot / 1000, published_curve, "-", color="C0", label="Published equation")
ax.plot(
mock_data["Electrolyte concentration [mol.m-3]"] / 1000,
mock_data["Electrolyte conductivity [S.m-1]"],
"s",
color="C1",
label="Mock data",
)
ax.plot(c_plot / 1000, fitted_curve, "--", color="C1", label="Fit to mock data")
ax.set_xlabel("LiPF$_6$ concentration [mol/L]")
ax.set_ylabel("Electrolyte conductivity [S/m]")
ax.set_title(f"Landesfeind 2019 EC:EMC (3:7) at T = {T_iso} K")
ax.legend()
fig.tight_layout()