2. Full-cell OCV¶
In this example, we use synthetic full-cell GITT data and known half-cell OCP parameters to determine the stoichiometry windows that give the correct full-cell OCV, a process often referred to as “electrode balancing”.
Before running this example, make sure to run the script true_parameters/generate_data.py to generate the synthetic data.
from pathlib import Path
import ionworkspipeline as iwp
import iwutil
from true_parameters.parameters import get_msmr_parameters
/Users/runner/work/ionworks-app/ionworks-app/.venv/lib/python3.12/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Cell balancing theory¶
Given the open-circuit potentials for each electrode, we combine them to get the full-cell open-circuit voltage
In this example, we fit the “lower excess capacity” and “upper excess capacity” instead of the stoichiometries at 0% and 100% state of charge. We use these names because the data typically does not reach the true 0% and 100% state of charge, due to kinetic limitations and/or electrolyte stability. The lower excess capacity is equal to \(q_{n}^\mathrm{0}\) in the negative electrode, and \(q_{p}^\mathrm{100}\) in the positive electrode. The upper excess capacity is equal to \(Q_{n} - q_{n}^\mathrm{100}\) in the negative electrode, and \(Q_{p} - q_{p}^\mathrm{0}\) in the positive electrode. \(Q\) is given by the actual total capacity observed in the experimental data (called “useable capacity” in the example).
Load the data¶
We load the synthetic GITT data and the extract the OCP from the relaxed voltages.
gitt_data = iwutil.read_df(Path("synthetic_data") / "full_cell" / "gitt.csv")
ocp_data = iwp.data_fits.preprocess.pulse_data_to_ocp(gitt_data)
Get the known parameters¶
In this example, we assume we already know the MSMR OCP parameters for the negative and positive electrodes. In practice, these parameters can be fitted using the half-cell OCV workflow.
msmr_params_n = get_msmr_parameters("negative")
msmr_params_p = get_msmr_parameters("positive")
known_params = {
**msmr_params_n,
**msmr_params_p,
"Ambient temperature [K]": 298.15,
}
Set up the parameters to fit¶
Now we set up our initial guesses for the parameters. As in the previous example, we create a dictionary of Parameter objects.
Q_use = ocp_data["Capacity [A.h]"].max()
Q_n_lowex = iwp.Parameter(
"Q_n_lowex", initial_value=0.01 * Q_use, bounds=(0, 0.2 * Q_use)
)
Q_p_lowex = iwp.Parameter("Q_p_lowex", initial_value=0.1 * Q_use, bounds=(0, Q_use))
Q_n_uppex = iwp.Parameter(
"Q_n_uppex", initial_value=0.1 * Q_use, bounds=(0, 0.2 * Q_use)
)
Q_p_uppex = iwp.Parameter("Q_p_uppex", initial_value=0.5 * Q_use, bounds=(0, Q_use))
parameters = {
"Negative electrode lower excess capacity [A.h]": Q_n_lowex,
"Positive electrode lower excess capacity [A.h]": Q_p_lowex,
"Negative electrode upper excess capacity [A.h]": Q_n_uppex,
"Positive electrode upper excess capacity [A.h]": Q_p_uppex,
"Negative electrode capacity [A.h]": Q_n_lowex + Q_use + Q_n_uppex,
"Positive electrode capacity [A.h]": Q_p_lowex + Q_use + Q_p_uppex,
"Usable capacity [A.h]": Q_use,
}
Run the fit¶
Now we are ready to run the fit. We create the objective, the DataFit object, and run the fit.
First, we create the MSMRFullCell objective function and pass in the data. We need to specify the voltage range of each electrode to use in the fit. The limits are used to evaluate the half-cell OCPs in the fit, and should therefore span a wider half-cell voltage range than we expect to see in the data.
Next, we create the DataFit object and pass in the objective and the parameters to fit. We can also customize the fitting process by passing in a cost function and selecting an optimizer.
# Create the objective
objective = iwp.objectives.MSMRFullCell(
ocp_data,
options={
# these are the voltage ranges over which to evaluate the half-cell OCP curves
"negative voltage limits": (0, 2),
"positive voltage limits": (2.5, 5),
},
)
# Create the DataFit object and run the fit
optimizer = iwp.optimizers.ScipyLeastSquares()
datafit = iwp.DataFit(objective, parameters=parameters, optimizer=optimizer)
results = datafit.run(known_params)
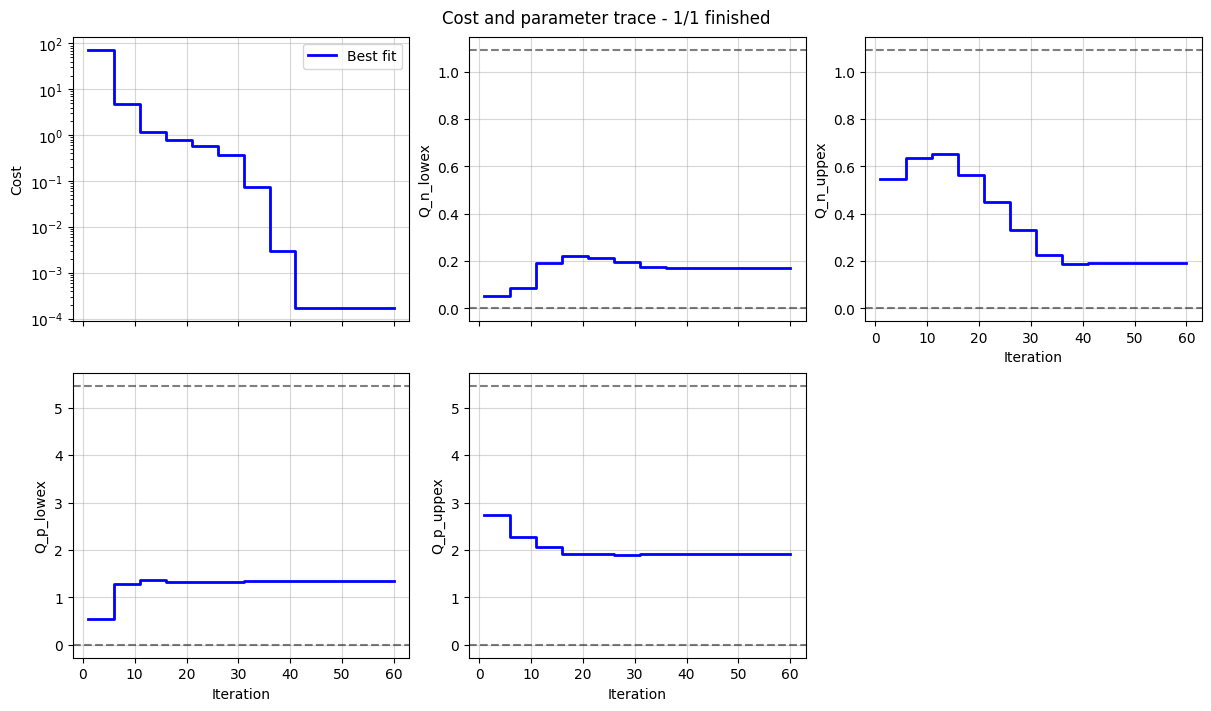
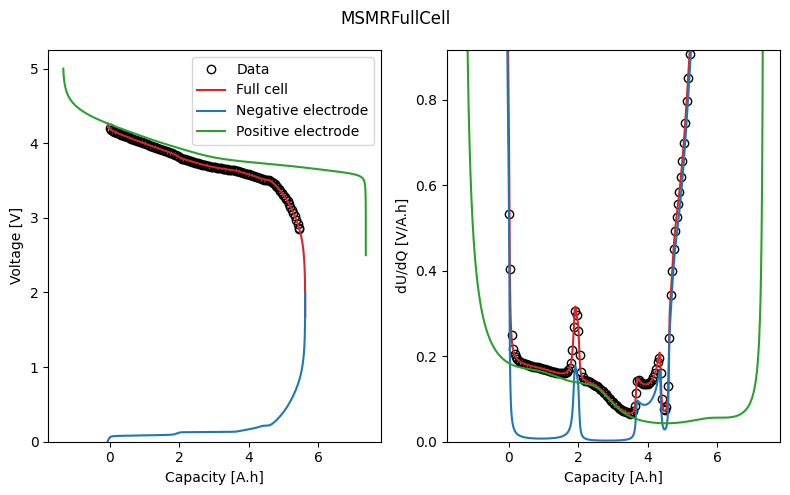
We can plot the trace of the cost function and parameter values during the fit, and the fitted OCV and dU/dQ curves. In the plot, we can see how the half-cell OCPs combine to give the full-cell OCV.
_ = datafit.plot_trace()
_ = datafit.plot_fit_results()
We can take a look at the fitted parameters by accessing the parameter_values attribute of the results object.
results.parameter_values
{'Negative electrode lower excess capacity [A.h]': 0.17027676100322262,
'Positive electrode lower excess capacity [A.h]': 1.3505682381729573,
'Negative electrode upper excess capacity [A.h]': 0.19320804419451476,
'Positive electrode upper excess capacity [A.h]': 1.9149720165918984,
'Negative electrode capacity [A.h]': 5.827850521509166,
'Positive electrode capacity [A.h]': 8.729905971076285,
'Usable capacity [A.h]': 5.464365716311429}